Our Lab invented molecular tension fluorescence microscopy
The first approach to map cell traction forces with pN sensitivity!
The cellular response to physical stimuli has been a topic of significant interest in cell biology that range from fundamental developmental biology to cancer diagnostics. For instance, stem cells have been shown to sense and respond to the stiffness of their underlying substrate, steering differentiation based on the mechanical properties of the cellular microenvironment.
External physical information is transmitted into the cells as the force or tension by cell membrane receptors. Our lab has focused on studying cellular tension at the molecular scale of receptor-ligand interaction and has pioneered this field by developing the first molecular tension probe (Nature Methods, 2011). This initial breakthrough has revolutionized mechanobiology study by providing tools to visualize molecular forces with the speed, convenience, and precision of conventional fluorescence microscopy.
The design of the tension probes is fairly simple and is akin to a macroscopic force gauge. Probes consist of a deformable linker (DNA, protein, or polymer) flanked by a fluorophore and a quencher. These probes are immobilized onto a substrate (glass, lipid bilayer, or hydrogels) and present a biological ligand that engages the receptor of interest. When a cell applies a specific pN forces to stretch the probe, the fluorophore is separated from the quencher, thus leading to an enhancement in signal (up to 100 fold enhancement).
INTEGRIN-MEDIATED MECHANOTRANSDUCTION
Integrins are adhesion receptors that span the plasma membrane and anchor cells to the external environment. We are interested in studying the interplay between mechanical forces and chemical signaling within integrins-mediated adhesions. Our initial work using MTFM showed that integrins can transmit forces>10 pN (JACS 2013). Subsequent work using MTFM probes tethered to patterned nanoparticles showed a relationship between ligand nanoscale spacing and force transmission (Nano Lett. 2014). Building on MTFM technology, we have since investigated processes such as cardiomyocyte differentiation, platelet activation, and inflammatory responses in asthma.
molecular force microscopy (MFM)
Compared with conventional traction force analysis methods, one drawback of molecular tension probe is lacking the information of force direction. To this end, we leveraged molecular tension probes and fluorescence polarization microscopy or structured illumination microscopy to measure the magnitude and 3D orientation of integrin forces. (Nat Methods, 2018 & Nat. Comm., 2021)
Super-resolution Imaging of Integrin-Mediated Cellular force
One of the advantages of molecular tension probes are their nature of detecting force at a single cell receptor-probe interaction. To achieve the higher resolution of cellular force measurement, we first integrated molecular tension probes with the DNA points accumulation for imaging in nanoscale topography (DNA-PAINT) technique. This tension-PAINT technique has achieved to map piconewton mechanical events with ~25-nm resolution. (Nat. Methods, 2020)
Single molecule tension imaging to study single-receptor force dynamics
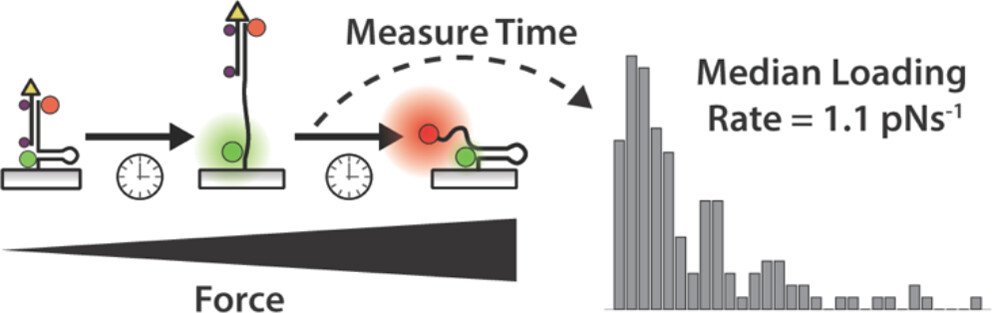
One outstanding question in mechanobiology pertains to the dynamics of forces transmitted by mechanoreceptors. How fast does a receptor ramp up mechanical load? How heterogeneous are force ramp rates? These are fundamental questions that remain unresolved and hamper our understanding of mechanotransduction. We addressed this question by integrating the single molecule imaging technique with the molecular tension probe containing two distinct force detection units. Sequential detection of two distinct mechanical events in single molecule resolution allowed one to calculate the force loading rate of integrin-ligand interaction under living cells’ focal adhesion. (JACS 2024)
Mechanically Induced Catalytic Amplification Reaction for Readout of Cellular Forces
Mechanics play a fundamental role in cell biology, but detecting piconewton (pN) forces is challenging because of a lack of accessible and high throughput assays. In this project, we are developing mechanically induced catalytic amplification reactions (MCR) for readout of cell forces. As a proof of concept, the assay was used to test the activity of a mechanomodulatory drugs and integrin adhesion receptor antibodies. To the best of our knowledge, this is the first example of a catalytic reaction triggered in response to molecular piconewton forces. The MCR may transform the field of mechanobiology by providing a new facile tool to detect receptor specific mechanics with the convenience of the polymerase chain reaction (PCR). (Nat. Biomed. Eng., 2023)
Force-induced Cell Tagging technique
Flow cytometry is used routinely to measure single-cell gene expression by staining cells with fluorescent antibodies and nucleic acids. In this project, we developed a new approach to tag mechanically active cells. We named the technique tension-activated cell tagging (TaCT) which labels cells fluorescently based on the magnitude of molecular force transmitted through cell adhesion receptors. As a proof-of-concept, we analyzed fibroblasts and mouse platelets after TaCT using conventional flow cytometry. (Nat. Methods, 2023)
Force-Selective Drug Delivery
The mechanical dysregulation of cells is associated with a number of disease states, that spans from fibrosis to tumorigenesis. Hence, it is highly desirable to develop strategies to deliver drugs based on the “mechanical phenotype” of a cell. To achieve this goal, we developed DNA mechanocapsules (DMCs) comprised of DNA tetrahedrons that are force responsive. DMCs are designed to encapsulate macromolecular cargos such as dextran and oligonucleotide drugs with minimal cargo leakage. We demonstrated force-induced mRNA knockdown of HIF-1α in a manner that is dependent on the magnitude of cellular traction forces. (Nat. Comms. 2024)
Future directions
Despite our contributions to over a decade of tool development in the area of molecular mechanobiology, we still see many questions that need to be answered. What is the relationship between molecular force and receptor conformation? Does the magnitude, orientation, and dynamics of force impact biochemical signaling? Are the measured forces on a substrate similar to those in vivo? How do different types of posttranslational modifications alter mechanobiology? What is the role of the plasma membrane in force transmission? As you see, we are still in the infancy of this field and the development of new tools is badly needed.
“Killer T cell attacking cancer” posted by Cambridge University, Feb 3, 2012. T cell shown in green, cancer cell shown in blue.
The immune system has a very difficult job. It has to find the metaphorical needle in the haystack and distinguish the very few infected and cancerous cells from the billions of normal cells throughout the body. To achieve this very difficult task, T cells, the workhorse cells for surveillance, have to physically crawl over and interact with target cells which in turn display snippets of their proteins on their plasma membrane. The recognition process is mediated by T cell receptors (TCRs) that signal once they bind antigens on target cells. TCR-rich microvilli push and pull on the apposing cell membrane as the T cell scans for antigens. This is clear from the video shown to the right. Videos like this prompted us to ask the following questions: Is the TCR-antigen complex experiencing pushing and pulling forces? What is the magnitude and duration of the force? How does force influence immune response? Many others in the field had postulated and demonstrated that the TCR-antigen bond is sensitive to force, but for us we were interested in measuring the intrinsic forces generated by the T cell and experienced by the antigen. We also wanted to understand the process of mechanosensing and mechanotransduction. Our approach has focused on developing tools to measure and in some cases control forces and then to record signaling outcomes. For reviews on this topic please see the following papers: DNA Nanotechnology for Investigating Mechanical Signaling in the Immune System (link), DNA Nanotechnology as an Emerging Tool to Study Mechanotransduction in Living Systems (link), and Mechanical Control of Antigen Detection and Discrimination by T and B Cell Receptors (link).
DNA-based MTFM to visualize TCR forces
Our work in this area started when we used molecular tension probes (see previous section on mechanobiology probes) to investigate TCR forces (PNAS 2016). We discovered that TCRs transmit piconewton-scale forces to their antigens (for CD8+ OT-1 cells 12pN>F>19pN). The force probe was comprised a force-responsive DNA hairpin structure flanked by a FRET pair which generates a fluorescent “on” signal upon force exertion. Our foray in this field was very exciting as we were able to demonstrate that the magnitude of these forces was highly dependent on kinase signaling and co-receptor engagement with the number of mechanical events detected increasing with antigen potency. Additionally, we found that these forces aid T cells in recognizing and initiating TCR stimulation upon engagement with their cognate antigens (PNAS 2016).
DNA tension probes for mechanical information storage
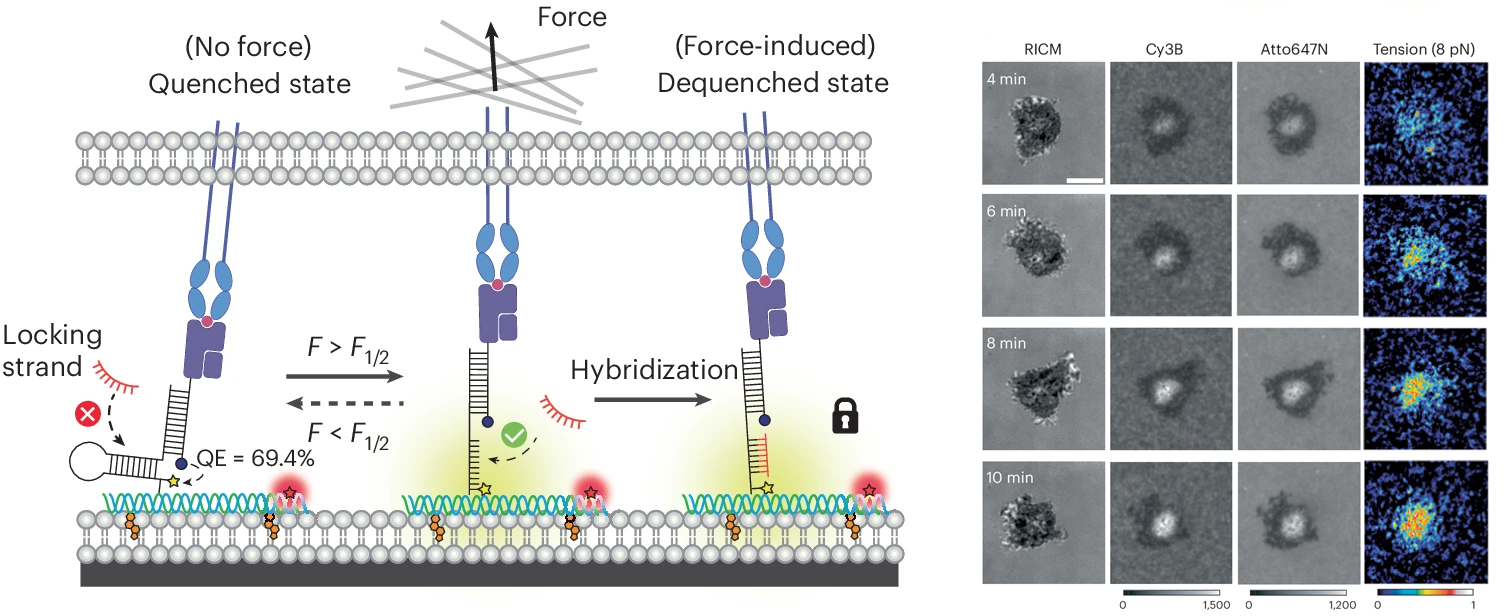
While we were able to visualize TCR-mediated forces with our initial DNA-based tension probe, often times the observed signal was relatively weak as we studied the interaction between TCRs and pMHCs, the natural ligand for the TCR. This is due to the transiency of this interaction; previous work has shown that most of these interactions have a bond liftetimes in the millisecond range. To capture these highly transient interactions, we developed a technique that allows for mechanically unfolded tension probes to be locked in their open state. This allows tension signal to accumulate over time and enables visualization of mechanical events that are not visible by observing a real time snapshot of a small population of unfolded probes (PNAS 2019). An important outcome of this study is the development of a new class of probes that trigger a chemical response (DNA binding) in response to a precise pN mechanical event. This is the basis for many subsequent technologies described below.
T cell co-receptor forces enhance T cell activation
In addition to the TCR-pMHC interaction, T cells engage a number of ligands on antigen presenting cells that promote either T cell activation or immune suppression. A key co-stimulatory interaction that promotes T cell activation by promoting cellular adhesion is the engagement of ICAM-1 by LFA-1 on the T cell surface. Using spectrally encoded DNA tension sensors, we visualized TCR and LFA-1 forces simultaneously as a T cell interacts with a substrate decorated with cognate pMHC and ICAM-1. Additionally, we discovered that tension between LFA-1 and ICAM-1 must be sustained to fully potentiate a T cell response. This was the first direct observation of LFA-1 exerting force onto ICAM-1 and analysis of the role that these forces play in T cell activation (Science Advances 2022).
TCR-pMHC serial mechanical engagement enhance T cell activation
We next became curious as to whether a single mechanical event is sufficient to trigger the TCR and mediate its response or whether the TCR needs repeated mechanical sampling of its antigen. In other words, does a single “handshake” provide the needed information to mount an immune response or would the TCR benefit from having multiple handshakes? This idea ties to the serial engagement model that proposes that a single antigen is sampled repeatedly by different TCRs to mediate an ultrasensitive response. So we were hoping to reveal the interplay between TCR mechanosensing and serial engagement. We designed a probe that “self-destructs” after a defined time following mechanical triggering. The FUSE probe recruits a nuclease enzyme only if mechanically triggered and the timing of the self-destruction is set by the enzyme concentration. This allowed us to study the relationship between TCR-antigen force duration and T cell stimulation and the results demonstrated that serial mechanical engagement enhances T cell activation. Therefore, it meant that the TCR takes advantage of repeated pN mechanical sampling to boost the levels of T cell stimulation. The work was the first demonstration of the potential cooperation between the mechanosensor and serial engagement models for T cell activation (JACS 2024).
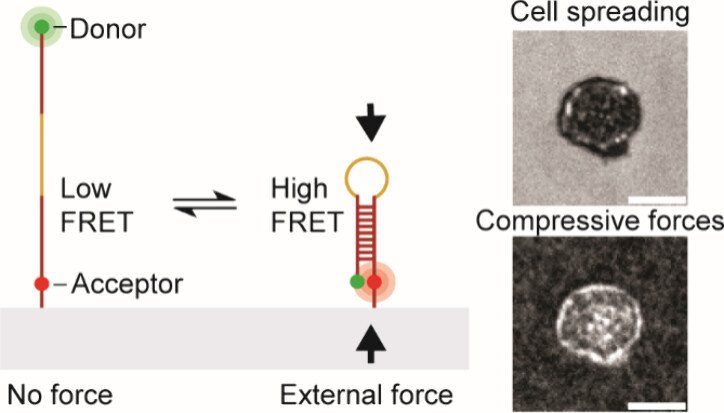
T cells exert compressive forces upon antigen engagement
While previous work robustly characterized tensile “pulling” forces between various T cell receptors and their ligands, we became curious as to whether the T cell “pushes” on the target cell. To address this question, we developed a molecular compression probe that allows compressive forces applied by T cells (and other cell types) to be visualized using FRET-based fluorescence microscopy. The key innovation here was to screen for DNA hairpins sequences that are pseudostable – at the verge of folding – but not fully folded at equilibrium. However, when an external compressive force is applied to the probe, then this encourages the hairpin to fold, and shifts the equilibrium toward the folded state. Effectively, this is akin to a crowding sensor. The probe generates high FRET signal only upon folding and the image to the right shows a map of T cell pushing forces applied to the substrate. This work demonstrates the first probe that maps molecular compressive forces and has been used to study molecular compressive forces exerted by T cells and platelets (JACS 2024).
DNA origami tension sensors to quantify forces at cell-cell interface
We next worked on finding methods of anchoring the DNA probes onto phospholipid membranes – which better mimic the physiological environment of TCR-antigen recognition. We particularly needed to keep probes at set distances (10-20 nms) away to avoid energy transfer processes probe-to-probe. We achieved this by creating DNA origami tension sensors (DOTS) that measure TCR-mediated forces in lipid membranes. DOTS can be tethered to virtually any lipid membrane, thus allowing one to study mechanotransduction at cell-cell junctions. DOTS are comprised of a rectangular nanosheet that presents a single force sensor on one face and cholesterol moieties for anchoring to membranes on the other face. We anchor DOTS to supported lipid membranes (SLBs) as well as live cell plasma membranes to present specific ligand and record the forces experienced by these ligands. The DOTS platform overcomes a long-standing limitation in this field, which is that lateral clustering of tension sensors leads to intermolecular FRET that creates artifacts and hinders quantitation of forces at fluid interfaces. We show unambiguously that T cells transmit >5 pN forces to their cognate antigens within live cell-cell junctions. We identify the source of TCR forces during initial antigen engagement and confirm that force is dependent on the nanometer height of the antigen and also on actin polymerization. We also demonstrate the potential for DOTS to be used in antigen screening by using spherical microparticles amenable to flow cytometry analysis of TCR-pMHC tension. This allows one to potentially quantify of hundreds of thousands T-cell mechanical properties within minutes and can be leveraged as a tool to screen antigen based on its potency to trigger biomechanical signaling (Nature Nanotech. 2024).
Future work
We are just starting to understand the role of molecular force in immune response and there remains a wide array of remaining questions. How does the cytoskeleton transmit force to the antigen? What is the precise relationship between force and bond lifetime? How are force magnitudes tuned? How does the structure of the TCR-antigen complex change with the application of pN forces? Does the co-receptor also experience forces? We recently showed that PD-1 and other co-receptors also transmit force. What are the role of these forces in co-receptors? And finally, as we “peel away the layers of the onion” and understand the molecular mechanisms of mechanotransduction in immune responses, we continually seeking to identify new diagnostics and therapeutics that can be realized by taking advantage of this new understanding.
RNA degraders pave the way to targeting the undruggable proteome
Unfortunately, 85% of the human proteome is considered undruggable because the protein structure is unknown, disordered, or lacking well-defined binding pockets. Many research groups have focused on developing RNA degraders to shift away from this inherent problem with the current protein-centric paradigm of medicinal chemistry. The most clinically successful approach to RNA degradation is with antisense oligonucleotides (ASOs). ASOs are small single-stranded DNA sequences that hybridize to a target mRNA via Watson-Crick-Franklin base pairing and induce the mRNA’s hydrolysis by the ubiquitous enzyme RNAseH. These therapeutics target a disease-associated protein at the transcript level rather than the protein level, enabling degradation of protein targets for which no small molecule inhibitor exists.
A major challenge of all therapeutics, however, including RNA degraders is selectively delivering and activating in target cells while leaving healthy cells alone. These challenges are particularly difficult for ASO development. Indeed, less than 1% of ASOs reach the cytoplasm due to their large molecular weights and the charged nature of the backbone. Moreover, ASOs can hybridize to partially complementary mRNAs causing off-target issues. The Salaita lab has been at the forefront of tackling the delivery, selectivity, and toxicity issues of ASOs by creating cutting-edge drug delivery vehicles, investigating methods to enhance sequence specificity using multivalency, and creating microRNA-responsive ASOs.
Densely functionalized nanodiscoidal nucleic acids potently degrade target mRNA
Delivery of nucleic acids can be hindered by nuclease susceptibility, endosome trapping, and clearance. Multiple nanotechnology scaffolds have offered promising solutions, and among these, lipid-based systems are advantageous because of their high biocompatibility and low toxicity. However, many lipid nanoparticle systems still have issues regarding stability, rapid clearance, and cargo leakage. We have designed lipid nanodisc vehicles that incorporate a modified Apo-A1 mimetic peptide to enable unprecedented DNA loading onto the disc scaffold, which we call nanodiscoidal nucleic acids (NNAs). We demonstrated that these NNAs functionalized with an ASO against HIF1-alpha potently reduce HIF1-alpha mRNA transcript levels compared to ASOs without a vehicle and demonstrated in 3D tumor spheroid models that these NNAs can penetrate into the necrotic core, a substantial challenge of conventional therapeutics. Thus these NNAs potentially offer an improved platform for ASO therapeutics. (ACS Chem. Biol. 2023)
pH-responsive nanoparticles as Trojan Horses to improve ASO drug release into the cytoplasm
One of the biggest challenges facing nucleic acid therapeutics is their inability to escape the endosome, the stomach of the cell. Indeed, 99% of DNA nanotherapeutics are estimated to be destroyed in the endosome before they can engage their target in the cytoplasm. To overcome this limitation, we have designed DNA Endosomal Escape Vehicle Response (DELVR). This new strategy enables DNA nanotherapeutics to escape the endosome by responding to the endosome's acidic environment using pH-responsive i-motif DNA and endosomal escape peptides. We have demonstrated that DELVR substantially increases drug potency compared to traditional nanotherapeutic vehicles and that DELVR mediates this increased potency via increased endosomal escape. Thus, the DELVR concept could offer a solution to the ongoing issue of poor cytosolic delivery of ASOs in viv. (ACS Nano 2024)
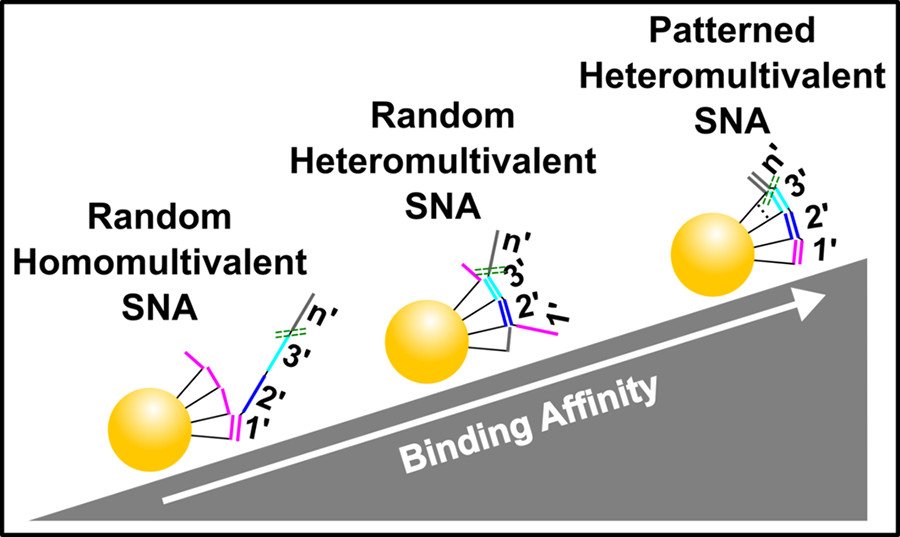
Heteromultivalency leads to enhanced target sequence binding avidity
Improving the affinity of nucleic acids to their complements is an important goal in antisense therapy and diagnostics. One potential approach to achieving this goal is to use multivalent binding, which often boosts the affinity between ligands and receptors, as exemplified by virus–cell binding. Inspired by nature, we have developed spatially patterned heteromultivalent DNA-nanoparticle conjugates and discovered that spatial patterning oligonucleotides on the surface of the nanoparticle drastically boosts target sequence binding avidity and affinity. These insights into DNA hybridization could lead to improvements in hybridization-based disease diagnostic probes and DNA-based therapeutics such as ASOs. (ACS Nano 2020)
Heteromultivalency leads to drastic improvements in binding specificity
Selectively binding to disease-associated single nucleotide polymorphisms (SNPs) is extremely challenging and detecting these mutations is necessary to prescribe effective cancer therapies, perform genetic analyses, and distinguish similar viral strains. Traditionally, SNP sensing uses short oligonucleotide probes that differentially bind the SNP and wild-type targets. However, DNA hybridization-based techniques require precise tuning of the probe’s binding affinity to manage the inherent trade-off between specificity and sensitivity. We have developed heteromultivalent nanoparticles that show extremely high specificity for SNPs and we demonstrated between heterozygous cis and trans mutations and between different strains of the SARS-CoV-2 virus. Our findings indicate that heteromultivalent hybridization offers substantial improvements over conventional monovalent hybridization-based methods. These insights into nucleic acid binding could lead to breakthroughs in SNP treatment and point-of-care diagnostics. (Nature Chem. 2023)
Conditional ASOs that respond to microRNA signals can differentiates between cell types and disease states
MicroRNA (miRNA) are regulatory RNAs that modulate gene expression and are implicated in a variety of cancers and other disease states. Indeed, some miRNAs are exclusively expressed in select tissues or are highly overexpressed (> 10 fold) in diseased tissue compared to healthy tissue. To leverage this tissue and disease-state specificity of miRNA, we have developed conditional ASOs, which can selectively activate in the presence of a trigger miRNA. These conditional ASOs use toehold-mediated strand displacement (TMSD) to peel off the ASO from the locking strand, freeing it up to hybridize to its mRNA target and thus induce its degradation. These results have broad implications in the field of precision medicine and the ASO field at large and could pave the way for next-generation therapeutics with improved specificity and limited systemic toxicity. (ACS Chem. Biol. 2021)
Future directions
Despite our over-decade-long contribution to the field of smart therapeutics, there are still many questions that need to be answered in order to fully harness the power of RNA degraders such as ASOs. For instance, is there a way to create biocompatible vehicles that deliver ASOs and other biologics to organs outside the liver, kidney, and spleen? Is the DELVR platform compatible with lipid nanoparticles and will this change in vehicle cause a change in ASO release? Are there better ways of functionalizing lipid nanoparticles for increased drug loading capacity? What is the fate and uptake mechanism of DNA-functionalized lipid nanodiscs? Would a lipid-based DELVR be well-tolerated in vivo? Would multivalency increase ASO specificity? Can TMSD be used in other ways to improve ASO specificity and potency? These questions and many others are in desperate need of answers, as they could hold the key to a brighter future for those suffering from diseases worldwide.
One feature common to all living systems is the requirement to generate mechanical forces that are fueled by chemical energy. For example, cells need to generate force to separate double stranded DNA to enable replication. Bacteria use ATP to power their flagella to move, and animals use ATP to drive muscle contraction and locomotion. Motor proteins, such as kinesin, dynein, and myosin, function by hydrolyzing ATP to change conformation and mediate movement. One grand challenge in the field is to develop synthetic versions of molecular motors and nanoscale machines. The motivation for making synthetic versions of motor proteins comes from anticipation that such devices could be used to power the next era of the industrial revolution – allowing society to make new classes of sensors, robots, computers and drugs with powerful capabilities. Indeed, this notion that we can create nanoscale devices and machines has inspired generations of fiction and non-fiction such as Eric Drexler’s Engines of Creation and Michael Crichton’s Prey.
Unfortunately, synthetic machines are still in their infancy. For example, we still don’t know how to miniaturize machines down to the molecular scale and still efficiently convert chemical energy to generate processive mechanical work. The motivation driving research under the DNA devices and motors subgroup is to address this grand challenge and to develop the basic blueprints for how to build synthetic machines that start to replicate the features of biological motors.
The following describes a brief chronological account of our contributions followed by a future outlook.
RNase H-powered Rolling motors
Our first “step” into this field was more of a “roll” with the development of the RNase H-powered high speed rolling motors that was reported in 2016. Most synthetic motor constructs worked by utilizing a bipedal – two legged - walker to move a motor along a DNA or RNA track. Leg binding induced hydrolysis of a “foothold” site on the track and this allowed for locomotion. The problem with this design was that motors frequently dissociated from the track because if both legs lost their footholds, then motors were lost to the bulk. Conversely, motors with many legs solved the dissociation problem but showed negligible speed because the leg movements were not coordinated. We solved the problem by 1) increasing leg polyvalency by orders of magnitude, 2) employing an enzyme with kcat orders of magnitude faster than the commonly used deoxyribozymes, and 3) patterning legs on a rigid spherical chassis that facilitates leg-to-leg coordination.
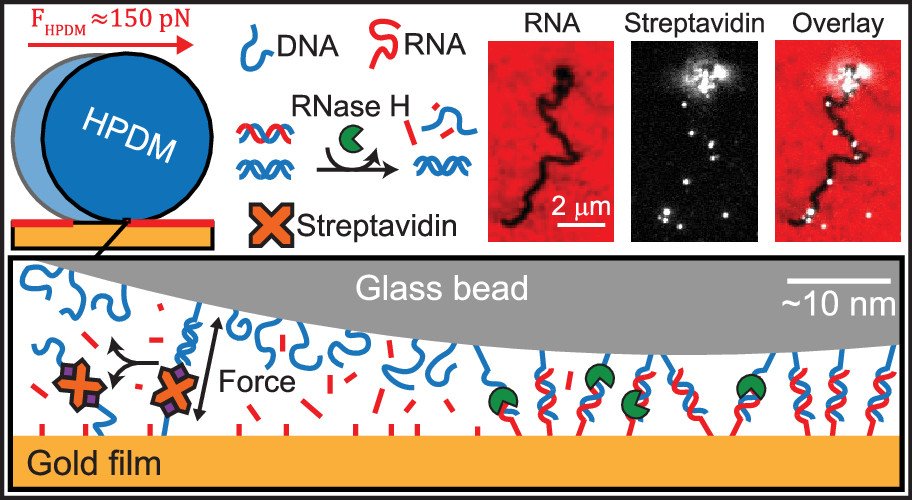
(Left) Motor chassis movement tracking in brightfield. (Right) Motor-localized fluorescently-tagged substrate depletion in TRITC.
Motors were comprised of micron scale silica spheres densely coated with DNA legs and allowed to bind a surface with a dense array of RNA foothold sites. When RNase H was added, motors moved at speeds of up to microns per minute and continued moving for millimeter distances. The video shows our rolling motors where the RNA footholds were tagged with a fluorescence dye. This first report showed that rolling motors exceeded the speed and processivity of all previously reported DNA walkers. Furthermore, we can monitor these motors in real time on a high-powered microscope, or even with a smartphone camera (Nature Nanotech. 2016)
Polyvalent DNA motors can generate 100+ pN of Force
We next asked “How much force is generated by rolling motors?”. To address this question, we created DNA-DNA tethers that acted as a molecular resistive tether that counteracted the force generated by the motors. We used DNA tethers because the forces needed to denature DNA duplexes was already well studied by the biophysics community. We titrated the density of DNA tethers so that we can investigate how many DNA duplexes are needed to overcome the force generated by motors and induce stalling (see figure below). Our work demonstrated that the motors rupture DNA duplexes in both the unzipping and shearing geometries (~12 and ~50pN), and even biotin-streptavidin bonds (~100-150 pN) which are often described as Nature’s strongest non-covalent bonds. Additionally, by utilizing Alexa647 (A647) fluorophore-functionalized streptavidin (SA) molecules this system became capable of molecular lithography. As when the biotin-streptavidin bonds rupture, the A647-SA is left on the substrate surface with the motor’s fluorescently-labelled substrate depletion track, thus depositing bright instances of “molecular ink” onto the a surface. (Nano letters 2019)
Nanoscale motors
We next wanted to investigate the basic rules and blueprints of how motor molecluar design influences its efficiency and activity. To achieve this goal, we created motors comprised of DNA origami which allows for highly precise (angstrom scale) molecular engineering of the chassis and DNA legs of motor. Motors were designed to resemble and “rolling pin” approximately 120 nm long and 20 nm wide. We found that these rod-shaped motors moved by rolling and hence translocated in a linear fashion (ballistically) on the RNA track. Importantly, the work demonstrated that the local DNA leg density and the rigidity of the chassis were critical in boosting motor efficiency. These parameters influence leg-to-leg coordination and rapid fuel consumption. (Angewandte Chemie 2020)
In a separate project, we created gold nanoparticle motors, and demonstrated that this design leads to enhanced speed/processivity. The gold particle “motor chassis” affords key advantages, which include ease of functionalization and display of a high density of DNA legs. In addition, gold particles strongly scatter visible light, providing unprecedented temporal (~50-500 msec) and spatial resolutions (~10 nm) for visualizing nanoscale dynamics. We used modeling and experiments to reveal that increasing leg DNA density boosts motor speed and processivity while DNA leg span only increases processivity. We show that motors translocate by rolling through a number of experiments including the observation that dimerized particles travel more ballistically. Interestingly, we discovered that motors exhibit bursts of high velocity punctuated with transient stalling – which is described as a Levy-type motion and is highly unusual for nanomaterials and typically observed in biological systems like migrating bacteria, birds, and marine predators. (ACS Nano 2022)
DNA computation systems with DNA motors
A major goal in the field is to create nanoscale robots that can deliver drugs, move cargo, and aid in manufacturing. Much of the progress toward DNA robotics has been enabled by nucleic acid reactions that demonstrate Boolean and fuzzy logic operations, which are the pillars of computation. However, to the best of our knowledge, there are no examples of DNA computing motors; i.e. DNA machines that consume chemical energy to produce mechanical work and that can sense and process information to guide mechanical activity. Creating genuine nanoscale robotic systems requires the integration of three components: a) chemical (input) sensing, b) information processing, and c) active mechanical function (output). Achieving this goal will also take us a step toward recapitulating basic features of single celled organisms that detect and process chemical information to guide directed motion. This has been a long-desired goal of the field, but there are major hurdles. For example, DNA computing systems are slow, and readout requires bulk fluorescence or single molecule AFM. It is not clear how one can link the input and output functions within a single machine. Indeed in a 2021 review on this topic, Chen and colleagues state that “A simpler and low-cost logic detection platform needs to be developed”.
We address this challenge by generating DNA motors that detect and process chemical inputs and transduce this information into easily detected macroscopic locomotion responses. Specifically, we create programmable DNA-based motors with onboard logic (DMOLs) that perform Boolean functions (NOT, YES, AND, and OR) with 15 min readout times. The logic gates were integrated into our rolling motors. The nucleic acid inputs triggered a binary DMOL response through toehold mediated strand displacement reactions, where 0 = no motion and 1= motion. Importantly, motor output transduces molecular information into macroscopic scale responses that can be detected using a conventional smart phone camera. Additionally, because DMOLs produce motion rather than color or fluorescence as the output, multiple unique DMOLs with different logic operations can be mixed on the same chip to process information in a massively parallelized fashion– and to scales that can never be achieved through fluorescence encoding. As a proof-of-concept of label-free encoding and multiplexing, we mixed three different DMOLs (OR, two-input AND, and three-input AND gates) detecting four nucleic acid inputs and showed that these can be read out in parallel using a smart phone microscope. (Nature Nanotech. 2022)
Time-lapse videos of Cy3 (red), Cy5 (blue) and FAM (green) fluorescence channels overlaid, acquired at 5 s intervals for a duration of 15 min
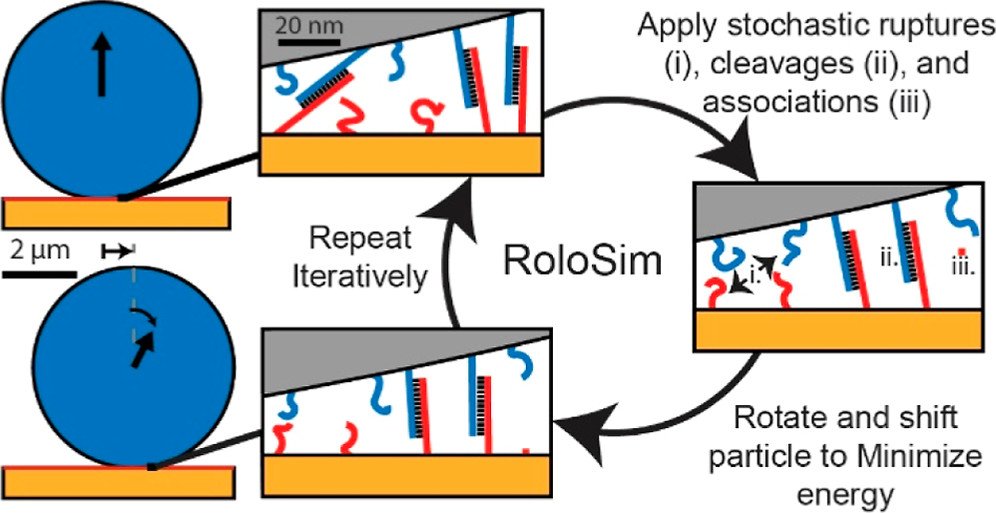
“RoloSim” modeling to predict motor performance
Next, we aimed to create computational models that can predict motor performance. Creating such models is important, as the number of potential parameters that one can tune are enormous and hence very difficulty to test out experimentally. To achieve this goal, we developed RoloSim, an adhesive dynamics software package for fine-grained simulations of motor translocation. RoloSim uses biophysical models for DNA duplex formation and dissociation kinetics to explicitly model tens of thousands of molecular scale interactions. These molecular interactions are then used to calculate the nano- and microscale motions of the motor. We used RoloSim to uncover how motor force and speed scale with several tunable motor properties such as motor size and DNA duplex length. Our results support our previously defined hypothesis that force scales linearly with polyvalency. We also demonstrate that motors can be steered with external force, and we provide design parameters for novel motor-based molecular sensor and nanomachine designs. (J. Phys. Chem. B 2022) Finally, RoloSim predicts the optimal parameters for creating better sensors and these predicts were subsequently validated in our “turbo-charged” motor work (see below).
“Turbo-charged” DNA motors
We next investigated how DNA leg sequence influences the speed and efficiency of motor translocation. DNA rolling motor speeds are largely determined by the interaction of RNase H with the DNA/RNA in the system, but simply increasing [enzyme] results in accelerated hydrolysis of fuel and reduced processivity. Instead, we investigated how DNA sequences influences the rate of DNA leg binding and dissociation from the fuel sites and ultimately tuning motion. By testing different DNA sequences in the substrate, we were able to identify sequences that create “turbo-charged” motors capable of a three-fold increase to instantaneous speed – reaching peak instantaneous velocities of 150 nm/sec. These turbo-charged motors were predicted to generate weaker forces, which should in principle enhance sensitivity when detect target molecules. Indeed, we observed that these motors could detect low copy numbers of DNA – and potentially demonstrate single molecule sensing by monitoring motor net displacements. (Angewandte Chemie 2024)
Timelapse video of the trajectories created by 0%, 33%, and 100% GC motors. The trajectories are centered at (0,0) and color-coded to indicate time. The trajectories were created using the x and y positions of the motors from brightfield particle tracking
Timelapse videos of Cy3 (red) fluorescence channels overlaid with brightfield acquired at 5 s intervals for a duration of 5 mins. The video was acquired ~30 mins after RNase H addition using a 100x 1.49 NA objective. On the left is the 33% GC motor and on the right is the 0% GC motor. The depletion tracks created by each motor are shown in black. Scale bar is 5 μm
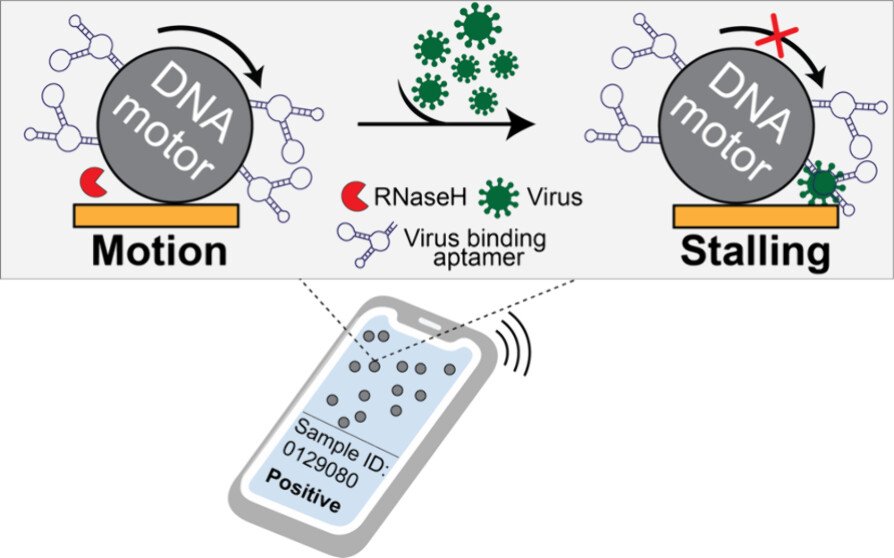
Mechanical virus detection using DNA motors
Our original rolling motor paper suggested that rolling motors could function as sensors of an analyte that mechanically resists motion. In other words, any molecule that stalls the motor could be detected. This is an exciting concept as it represents a fundamentally different approach to analytical chemistry that deviates from the conventional “bind, wash, and transduce” paradigm. Because we started this work during COVID, we decided to focus on detection of viral particles. We termed this strategy as RoloSense, which involves using motor movement to detect the presence of viruses. Here we display virus-specific DNA aptamers on both the motor and the surface. Viruses physically bridge the motor and the surface which induces stalling. Motors perform a “mechanical test” of the viral target and stall in the presence of whole virions, which represents a unique mechanism of transduction distinct from conventional assays. Rolosense can detect SARS-CoV-2 spiked in artificial saliva and exhaled breath condensate with a sensitivity of 10^3 copies/mL and discriminates among other respiratory viruses. The assay is modular and amenable to multiplexing. As a proof of concept, we show that readout can be achieved using a smartphone camera with a microscopic attachment in as little as 15 min without amplification reactions. These results show that mechanical detection using Rolosense can be broadly applied to any viral target and has the potential to enable rapid, low-cost point-of-care screening of circulating viruses. (ACS. Cent. Sci 2024)